Speed measurement
Here we will compare the speed of plotting UnfoldMakie with MNE (Python) and EEGLAB (MATLAB).
Three cases are measured:
- Single topoplot
- Topoplot series with 50 topoplots
- Topoplott animation with 50 timestamps
Note that the results of benchmarking on your computer and on Github may differ.
using UnfoldMakie
using TopoPlots
using BenchmarkTools
using Observables
using CairoMakie
using Statistics
using PythonPlot;
using PyMNE; CondaPkg Found dependencies: /home/runner/.julia/packages/CondaPkg/8GjrP/CondaPkg.toml
CondaPkg Found dependencies: /home/runner/.julia/packages/PyMNE/AlJE6/CondaPkg.toml
CondaPkg Found dependencies: /home/runner/.julia/packages/PythonCall/83z4q/CondaPkg.toml
CondaPkg Found dependencies: /home/runner/.julia/packages/PythonPlot/oS8x4/CondaPkg.toml
CondaPkg Dependencies already up to dateData input
dat, positions = TopoPlots.example_data()
df = UnfoldMakie.eeg_array_to_dataframe(dat[:, :, 1], string.(1:length(positions)));Topoplots
UnfoldMakie.jl
@benchmark plot_topoplot(dat[:, 320, 1]; positions = positions)BenchmarkTools.Trial: 71 samples with 1 evaluation per sample.
Range (min … max): 46.941 ms … 1.319 s ┊ GC (min … max): 0.00% … 95.96%
Time (median): 49.811 ms ┊ GC (median): 0.00%
Time (mean ± σ): 75.526 ms ± 162.277 ms ┊ GC (mean ± σ): 33.32% ± 15.51%
█
█▆▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▄ ▁
46.9 ms Histogram: log(frequency) by time 578 ms <
Memory estimate: 11.81 MiB, allocs estimate: 194551.UnfoldMakie.jl with DelaunayMesh
@benchmark plot_topoplot(
dat[:, 320, 1];
positions = positions,
topo_interpolation = (; interpolation = DelaunayMesh()),
)BenchmarkTools.Trial: 56 samples with 1 evaluation per sample.
Range (min … max): 57.024 ms … 1.581 s ┊ GC (min … max): 0.00% … 95.81%
Time (median): 62.555 ms ┊ GC (median): 0.00%
Time (mean ± σ): 89.585 ms ± 202.883 ms ┊ GC (mean ± σ): 30.19% ± 12.80%
▄▁▁▁▄ ▁▁▁ █▄ ▄▁▄ ▁
▆▁▁▁▆▁▁▁▁▆▁▁▆▁▁▁▁▆▁▆█████▆▆███▆██▁▆▆███▁▁▆█▆▆▆▁▆▆▁▆▁▆▁▁▁▁▁▁▆ ▁
57 ms Histogram: frequency by time 67.6 ms <
Memory estimate: 11.81 MiB, allocs estimate: 194558.MNE
posmat = collect(reduce(hcat, [[p[1], p[2]] for p in positions])')
pypos = Py(posmat).to_numpy()
pydat = Py(dat[:, 320, 1])
@benchmark begin
f = PythonPlot.figure()
PyMNE.viz.plot_topomap(
pydat,
pypos,
sphere = 1.1,
extrapolate = "box",
cmap = "RdBu_r",
sensors = false,
contours = 6,
)
f.show()
endBenchmarkTools.Trial: 320 samples with 1 evaluation per sample.
Range (min … max): 13.464 ms … 213.789 ms ┊ GC (min … max): 0.00% … 0.00%
Time (median): 14.166 ms ┊ GC (median): 0.00%
Time (mean ± σ): 15.642 ms ± 15.966 ms ┊ GC (mean ± σ): 0.00% ± 0.00%
▂▅▇█▇▅
███████▁▅▅▁▄▄▁▁▁▁▁▁▁▁▁▁▁▁▁▁▄▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▄ ▆
13.5 ms Histogram: log(frequency) by time 25.4 ms <
Memory estimate: 3.30 KiB, allocs estimate: 98.Topoplot series
Note that UnfoldMakie and MNE have different defaults for displaying topoplot series. UnfoldMakie in plot_topoplot averages over time samples. MNE in plot_topopmap displays single samples without averaging.
UnfoldMakie.jl
@benchmark begin
plot_topoplotseries(
df;
bin_num = 50,
positions = positions,
axis = (; xlabel = "Time windows [s]"),
)
endBenchmarkTools.Trial: 2 samples with 1 evaluation per sample.
Range (min … max): 2.363 s … 2.799 s ┊ GC (min … max): 0.00% … 11.16%
Time (median): 2.581 s ┊ GC (median): 6.05%
Time (mean ± σ): 2.581 s ± 308.237 ms ┊ GC (mean ± σ): 6.05% ± 7.89%
█ █
█▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁█ ▁
2.36 s Histogram: frequency by time 2.8 s <
Memory estimate: 351.58 MiB, allocs estimate: 5076848.MNE
easycap_montage = PyMNE.channels.make_standard_montage("standard_1020")
ch_names = pyconvert(Vector{String}, easycap_montage.ch_names)[1:64]
info = PyMNE.create_info(PyList(ch_names), ch_types = "eeg", sfreq = 1)
info.set_montage(easycap_montage)
simulated_epochs = PyMNE.EvokedArray(Py(dat[:, :, 1]), info)
@benchmark simulated_epochs.plot_topomap(1:50)BenchmarkTools.Trial: 7 samples with 1 evaluation per sample.
Range (min … max): 753.465 ms … 772.882 ms ┊ GC (min … max): 0.00% … 0.00%
Time (median): 764.776 ms ┊ GC (median): 0.00%
Time (mean ± σ): 763.952 ms ± 8.257 ms ┊ GC (mean ± σ): 0.00% ± 0.00%
▁ ▁ ▁ ▁ ▁ █
█▁▁▁▁▁▁▁█▁▁▁█▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁█▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁█▁▁▁▁▁▁▁█ ▁
753 ms Histogram: frequency by time 773 ms <
Memory estimate: 2.36 KiB, allocs estimate: 68.MATLAB
Running MATLAB on a GitHub Action is not easy. So we benchmarked three consecutive executions (on a screenshot) on a server with an AMD EPYC 7452 32-core processor. Note that Github and the server we used for MATLAB benchmarking are two different computers, which can give different timing results.
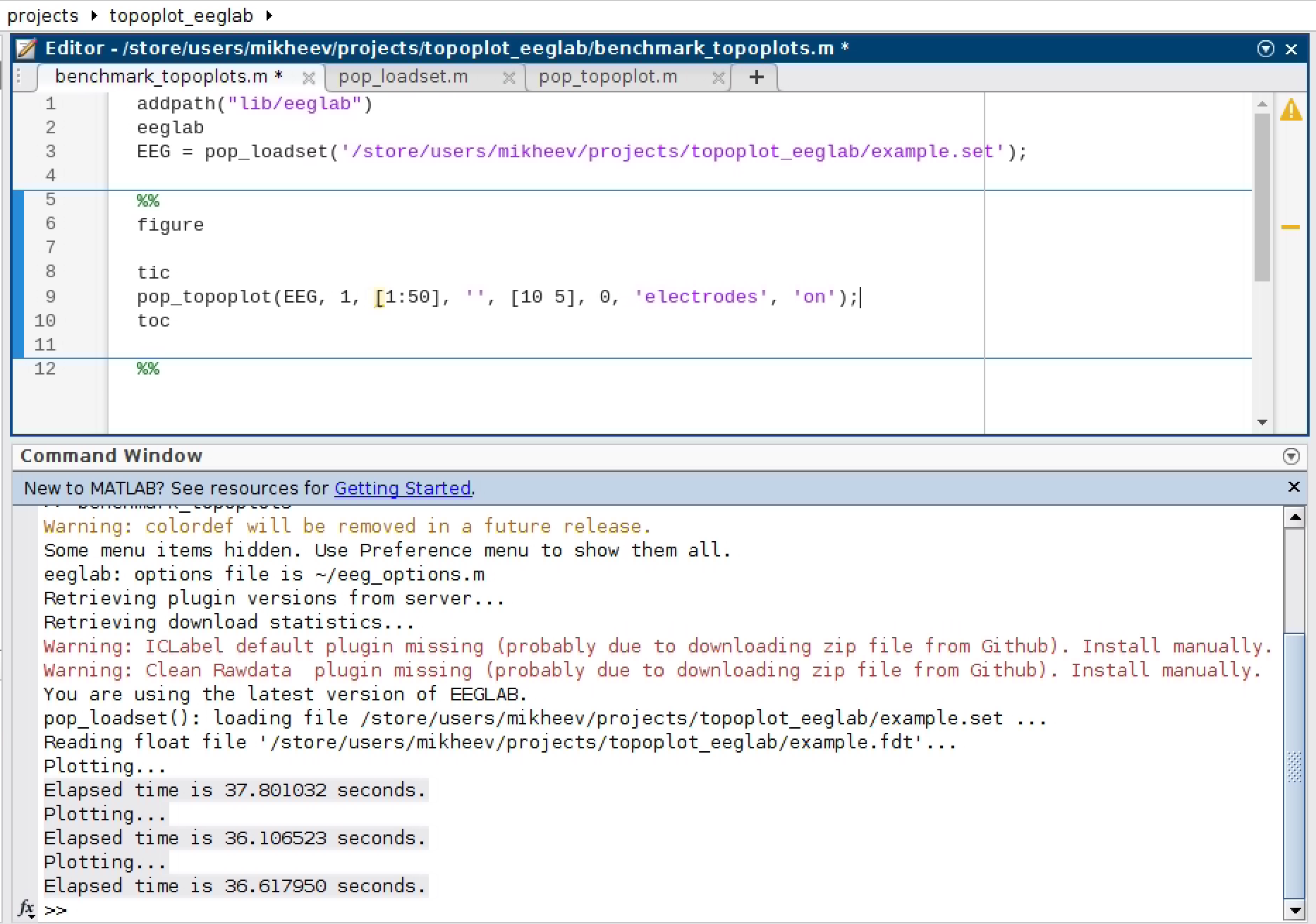
Animation
The main advantage of Julia is the speed with which the figures are updated.
timestamps = range(1, 50, step = 1)
framerate = 5050UnfoldMakie with .gif
vals = vec(dat[:, :, 1])
p01, p99 = quantile(vals, [0.01, 0.99])
m = max(abs(p01), abs(p99))
cr = Float32.((-m, m))
@benchmark begin
f = Makie.Figure()
dat_obs = Observable(dat[:, 1, 1])
plot_topoplot!(f[1, 1], dat_obs, positions = positions, visual = (; contours = false, colorrange = cr),)
record(f, "topoplot_animation_UM.gif", timestamps; framerate = framerate) do t
dat_obs[] = @view(dat[:, t, 1])
end
endBenchmarkTools.Trial: 3 samples with 1 evaluation per sample.
Range (min … max): 2.045 s … 2.229 s ┊ GC (min … max): 0.00% … 1.97%
Time (median): 2.182 s ┊ GC (median): 1.84%
Time (mean ± σ): 2.152 s ± 95.753 ms ┊ GC (mean ± σ): 1.30% ± 1.10%
█ █ █
█▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁▁█▁▁▁▁▁▁▁▁▁▁▁▁▁█ ▁
2.04 s Histogram: frequency by time 2.23 s <
Memory estimate: 120.33 MiB, allocs estimate: 416721.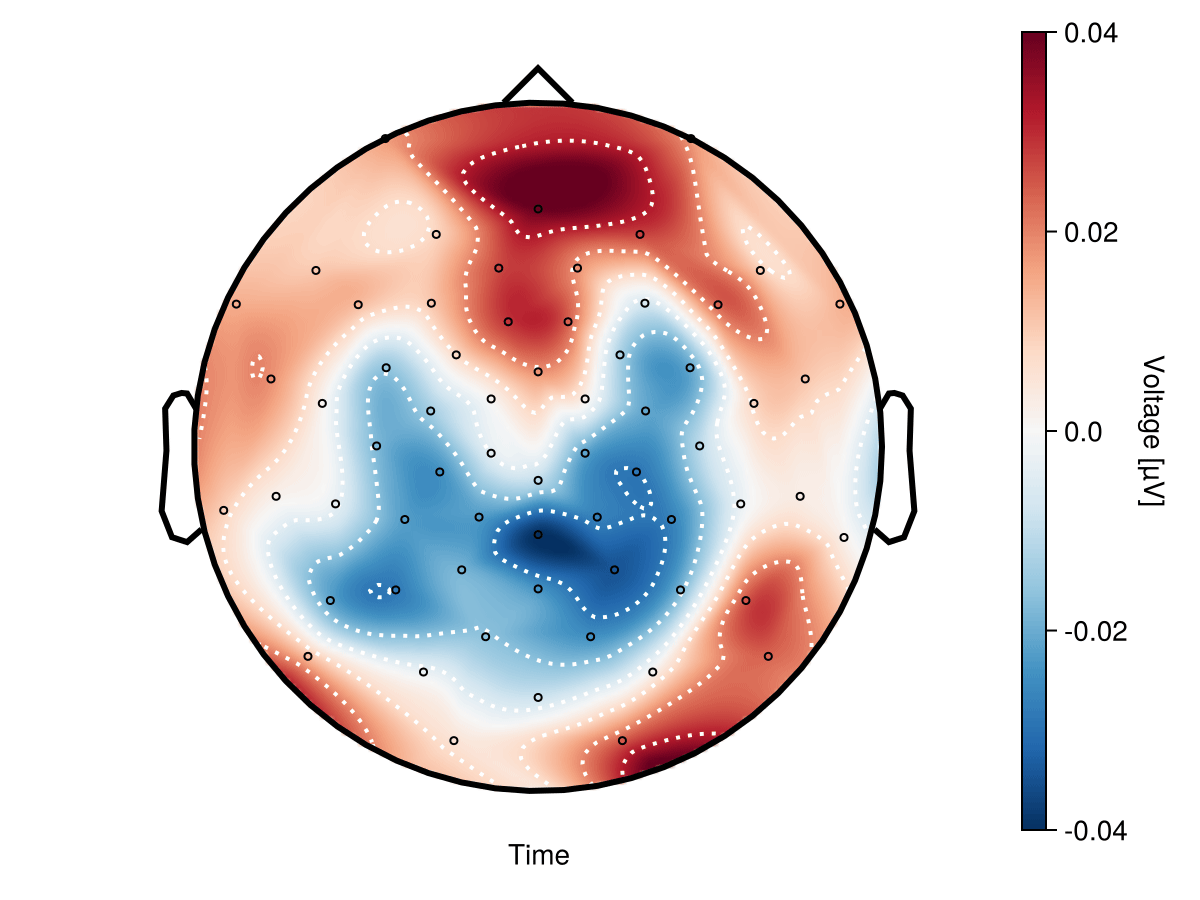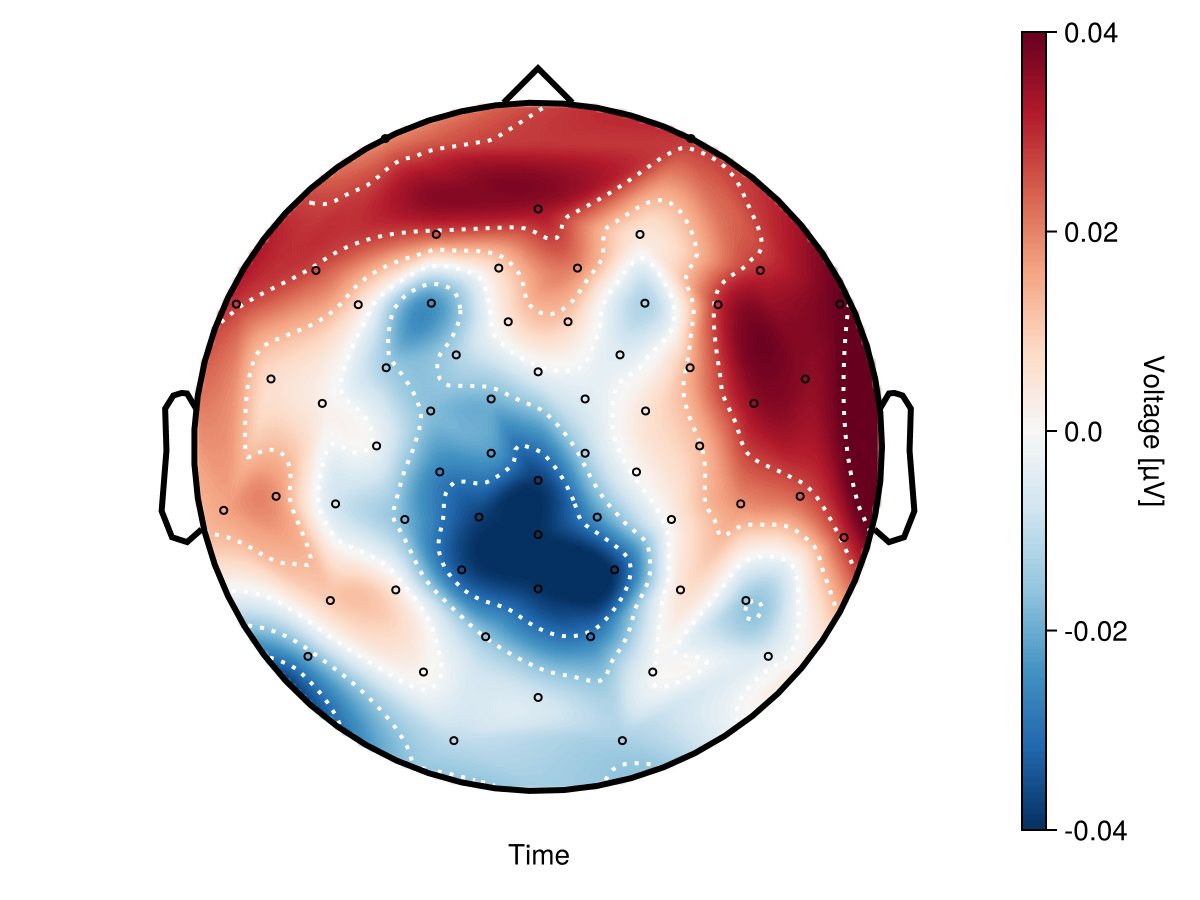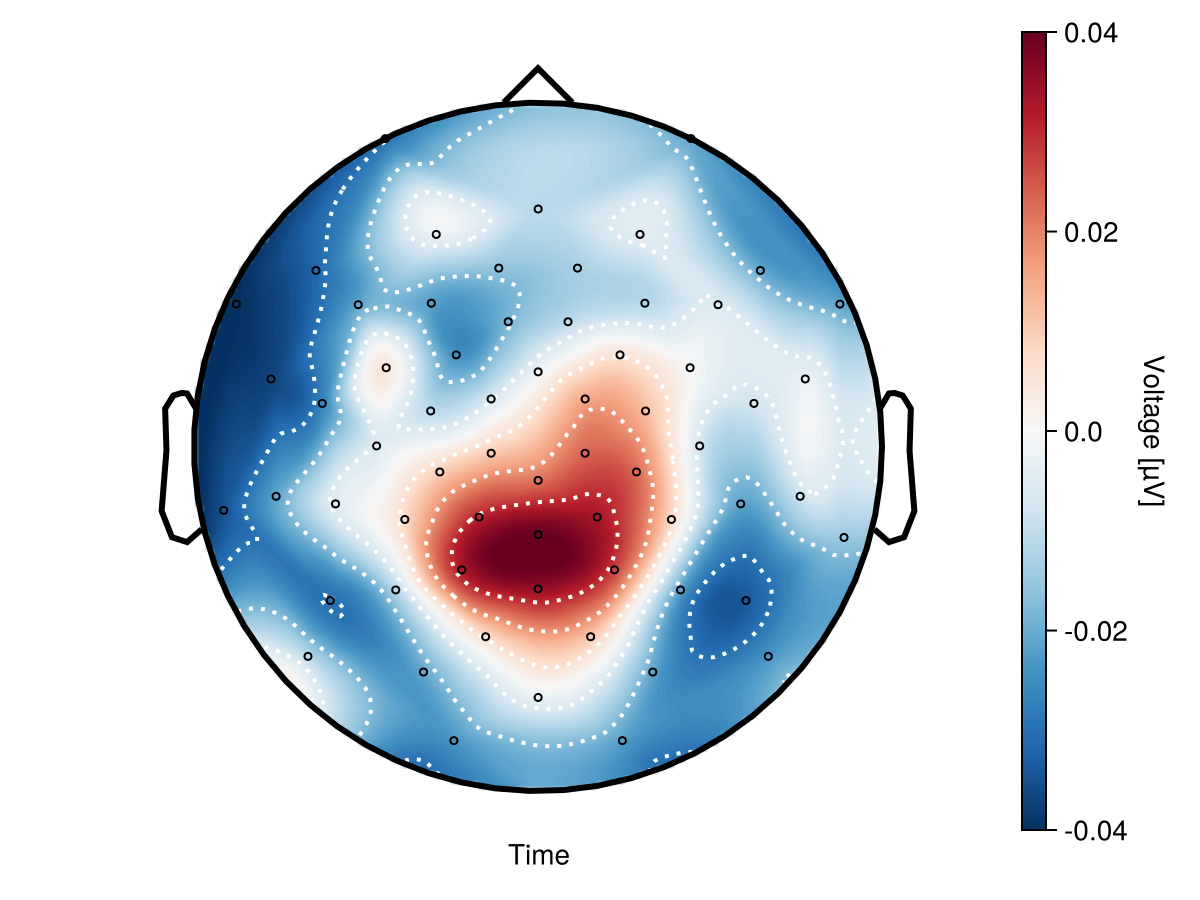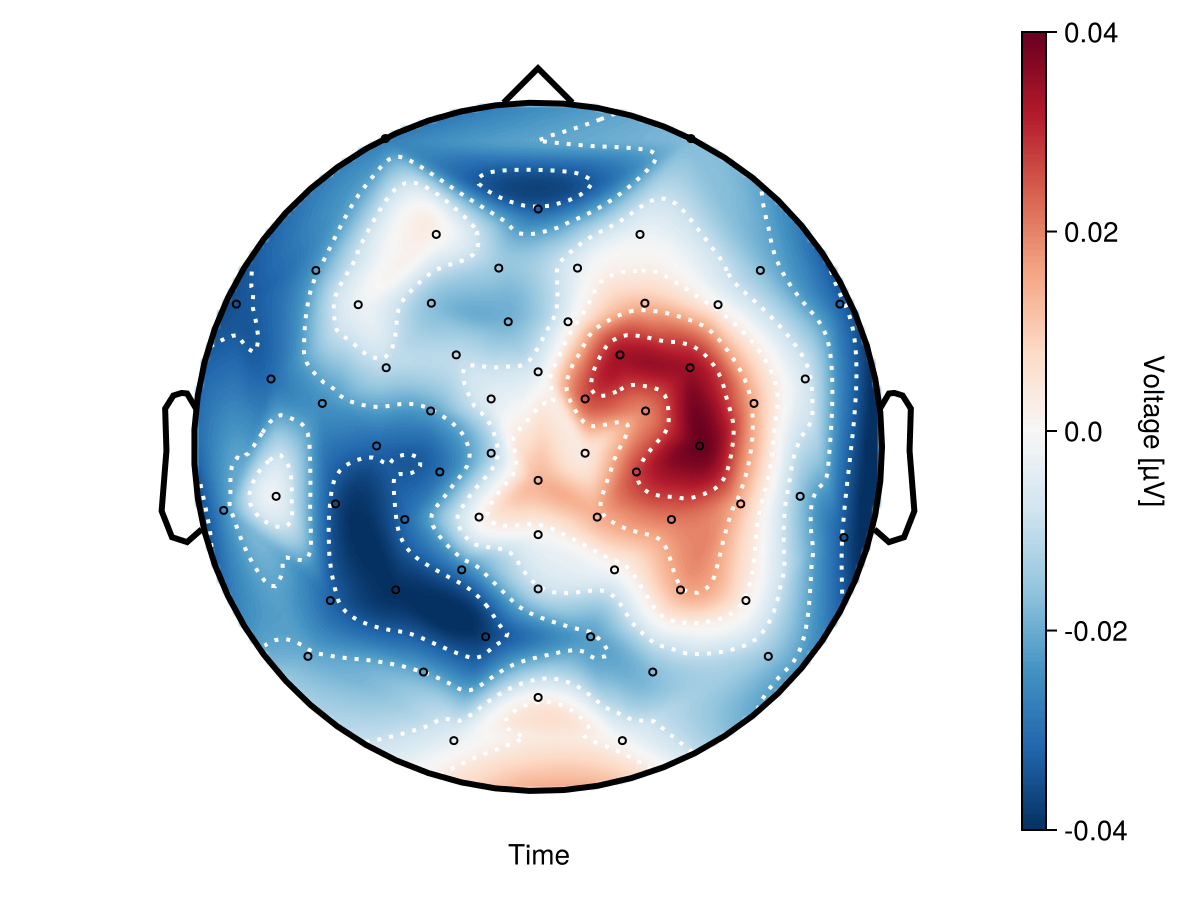
MNE with .gif
@benchmark begin
fig, anim = simulated_epochs.animate_topomap(
times = Py(timestamps),
frame_rate = framerate,
blit = false,
image_interp = "cubic", # same as CloughTocher
)
anim.save("topomap_animation_mne.gif", writer = "ffmpeg", fps = framerate)
endBenchmarkTools.Trial: 1 sample with 1 evaluation per sample.
Single result which took 8.790 s (0.00% GC) to evaluate,
with a memory estimate of 3.03 KiB, over 96 allocations.Note, that due to some bugs in (probably) PythonCall topoplot is black and white.
This page was generated using Literate.jl.